Contents
Exercise: RNA miniprep
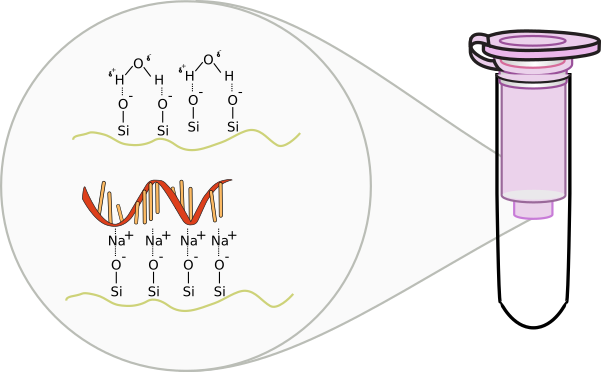
RNA purification occurs similar to DNA preparations. A silica based column is used where DNA is excluded from binding based on size and through an additional DNA digestion step using the enzyme DNase I. RNA is extremely fragile and prone to degradation. Because of this, separate pipettes and plastics are usually used in labs to reduce the amount of exposure to environmental or experimental RNase. When handling RNA, be extremely careful of contaminating the buffers or samples. Always wear gloves as skin carries RNase enzymes. Refrain from talking as to not contaminate the area with RNase found in saliva.
RLT = RNA lysis buffer: contains guanidine, a harsh denaturant
RW1 = RNA wash buffer
RPE = Second RNA wash buffer with Ethanol
RDD = DNase digestion buffer
- Harvest a maximum of 1 x 107 cells, as a cell pellet or by direct lysis in the vessel. Add the appropriate volume of Buffer RLT and vortex vigorously.
- If < 5 x 106 cells → 350 μl RLT (< 6cm plate)
- if ≤ 1 x 107 cells → 600 μl RLT (6-10cm plate)
- Add 1 volume of 70% ethanol to the lysate, and mix well by pipetting. Do not centrifuge. Proceed immediately to next step.
- Transfer up to 700 μl of the sample, including any precipitate, to an spin column placed in a 2 ml collection tube (supplied).
- Close the lid, and centrifuge for 15 s at ≥8000 x g.
- Discard the flow-through.
- Wash: Add 350 μl Buffer RW1 to spin column, close lid, centrifuge for 15 s at ≥8000 x g (≥10,000 rpm). Discard flow-through.
- Add 10 μl DNase I stock solution (see above) to 70 μl Buffer RDD. Mix by gently inverting the tube
- Remove DNA (optional): Add DNase I incubation mix (70 μl) directly to spin column membrane, and place on benchtop (20–30°C) for 15 min.
- Wash: Add 350 μl Buffer RW1 to spin column, close lid, centrifuge for 15 s at ≥8000 x g. Discard flow-through.
- Add 700 μl Buffer RW1 to the spin column. Close the lid, and centrifuge for 15 s at ≥8000 x g. Discard the flow-through.
- Add 500 μl Buffer RPE to the spin column. Close the lid, and centrifuge for 15 s at ≥8000 x g. Discard the flow-through.
- Add 500 μl Buffer RPE to the spin column. Close the lid, and centrifuge for 2 min at ≥8000 x g.
- Discard all flow-through and centrifuge at full speed for 1 min to dry the membrane.
- Place the spin column in a new 1.5 ml collection tube. Add 30 μl RNase-free water directly to the spin column membrane.
- Close the lid, and centrifuge for 1 min at ≥8000 x g to elute the RNA.
- Add 30 μl RNase-free water directly to the spin column membrane. Close the lid, and centrifuge for 1 min at ≥8000 x g to elute the RNA.
Reverse Transcription
The Central Dogma of Molecular Biology was proposed by Francis Crick, the co-describer of the double stranded helical structure of DNA. This “dogma” was a statement to describe the flow of genetic information to show that DNA houses or stores data that is transcribed into RNA that is subsequently translated from nucleotides into amino acids through the machinery of the ribosomes. Since DNA is relatively static in it’s ability to store genetic information, the expression of this stored data into the intermediate RNA or to the final protein product is of great significance. Imagine that the DNA in the nucleus of your cheek cells is identical to the DNA of the nucleus of cells in your liver. While the instructions are identical, these are clearly different cells that have a difference in expression of proteins. Imagine a hard drive on a computer that stores information as 1’s and 0’s. These 1’s and 0’s do not have meaning until specific programs are called upon to act on this information. Likewise, different programs are called to use the instructions of your DNA to make a cheek cell different than a liver cell.
In 1970, Howard Temin and David Baltimore independently isolated an enzyme from the Rous Sarcoma Virus and Murine Leukemia Virus, respectively. This enzyme was capable of violating the Central Dogma. The genomes of these viruses consist of RNA, not DNA. During the infection process, this enzyme is responsible converting the RNA into DNA in a process called reverse transcription. This enzyme is logically called reverse transcriptase (RT). This discovery was rewarded with Nobel Prize in 1975. Later on, more viruses were discovered that were composed of RNA genomes that utilized this process, including HIV. Other enzymes within cells were also recognized to have reverse transcriptase activity, such as telomerase and retrotransposases. In molecular biology, these enzymes are used to convert mRNA into complementary copies of DNA called cDNA. The sum total of everything that is transcribed into RNA is referred to as the transcriptome. Synthesis of cDNA from any transcribed RNA can then be used for transcriptome analysis.

Exercise: Reverse Transcription of Eukaryotic mRNA
mRNA from eukaryotes are modified with 3′ polyadenylated tails. Oligo-dT primers can be used to prime the reverse transcription process of all mRNAs. All solutions should be kept on ice.
- Determine the concentration of total RNA
- Adjust concentration of RNA to 0.1mg/ml using Rnase free water
- Combine 10 ml RNA, 1 ml Oligo-dT (50μM), and 1 ml dNTP Mix (10 mM each)
- Denature mixture at 65°C for 5 minutes and then place on ice
- Combine the following in a separate tube
- 4μl Buffer 5X → contains all salts and pH buffer
- 2μl 0.1 M DTT → a reducing agent to mimic the cellular environment
- 1μl RNaseOUT (40U/μl) → an RnaseA inhibitor
- 1μl SuperScript III RT → the reverse transcriptase enzyme
- After the denatured mixture has been sufficiently cooled, ad 8μl enzyme mixture
- incubate 45°C for 1 hour
- deactivate enzyme by incubating 75°C for 10 minutes
- Store your cDNA in the freezer
Tags: integration of knowledge


