- Outline the primary techniques utilized in recombinant DNA experiments.
- Summarize the problems in isolating, identifying, and cloning a single gene.
- Explain the action of restriction endonucleases and DNA ligases, and describe how each is used in recombinant DNA technology.
- Explain what DNA clones are, and describe 3 techniques commonly used to obtain them.
- Identify the role of vectors in recombinant DNA technology and give several specific examples of such vectors.
- Explain the special measures that have been used to introduce genes experimentally into plant and animal cells.
- Define restriction-fragment-length polymorphisms (RFLPs), and give one example of the way RFLPs are used to diagnose human genetic disorders.
- Explain the purpose and scope of the human genome project.
- List at least 5 potential or realized applications of recombinant DNA technology.
- Summarize the concerns that have been named about recombinant DNA research and the policies that have been formulated to address these concerns.
Contents
Analyzing DNA
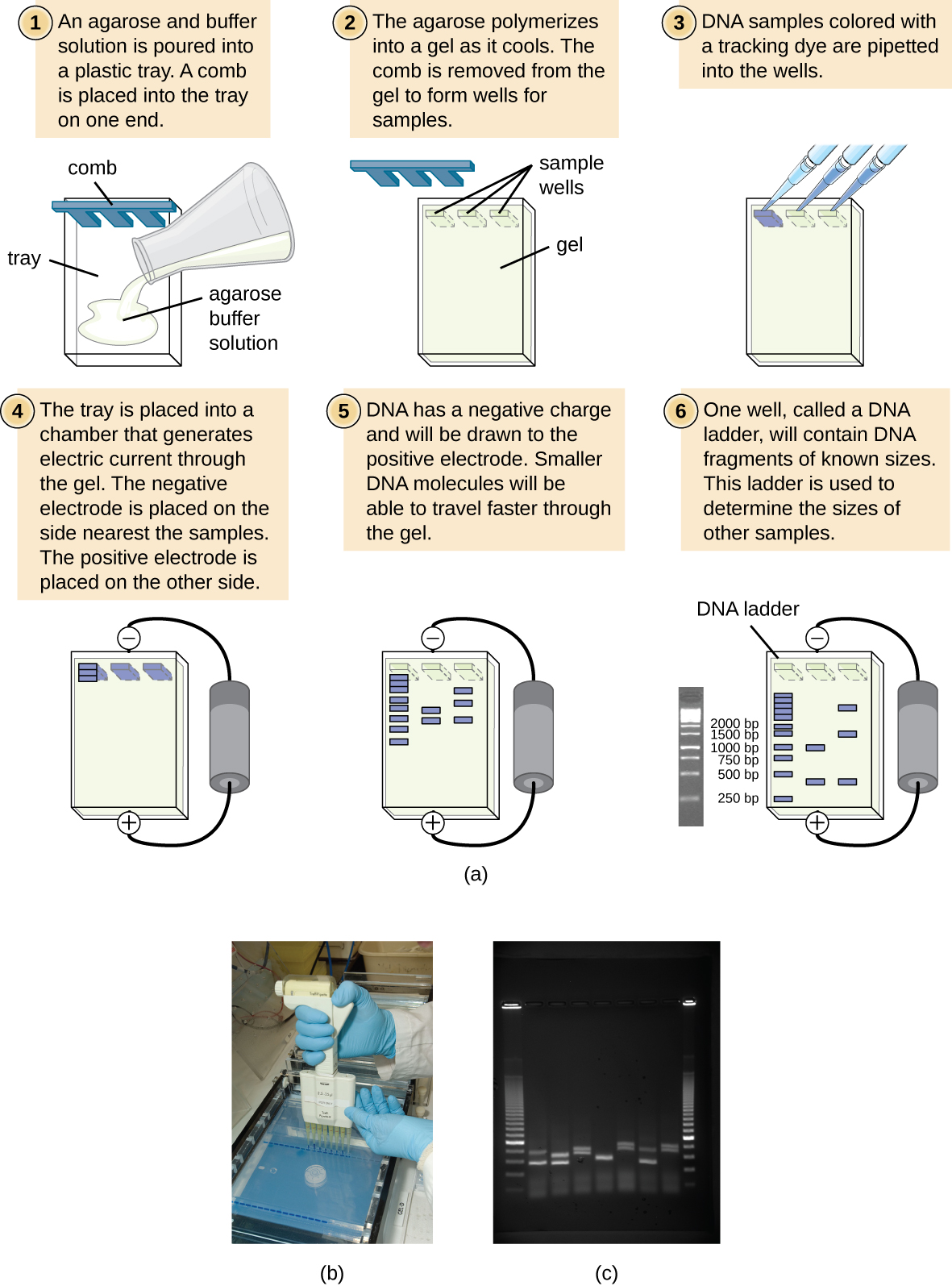
Agarose Gel Electrophoresis

Agarose is a linear carbohydrate polymer purified from the cell walls of certain species of algae. Agar is a combination of the crude extract that contains agarose and the smaller polysaccharide agaropectin. When dissolved and melted in liquid, agarose strands become tangled together to form a netting that holds the fluid in a gel. Reduction of the fluid creates a higher percentage gel that is firmer and contains smaller pores within the netting.
 Placing a comb within the melted agarose creates spaces that allow for the insertion of samples when the gel is solidified. Molecules can traverse through the pores as they are drawn by electrical currents. Charged compounds will migrate towards the electrode of opposite charge but migration rate will be influenced by the size of the molecules. Smaller compounds can easily traverse through the webbing while larger items are retarded by the pore size. Follow this simulation to get a better idea of how we use Agarose Gel Electrophoresis in molecular biology to study DNA fragments.
Placing a comb within the melted agarose creates spaces that allow for the insertion of samples when the gel is solidified. Molecules can traverse through the pores as they are drawn by electrical currents. Charged compounds will migrate towards the electrode of opposite charge but migration rate will be influenced by the size of the molecules. Smaller compounds can easily traverse through the webbing while larger items are retarded by the pore size. Follow this simulation to get a better idea of how we use Agarose Gel Electrophoresis in molecular biology to study DNA fragments.
DNA molecules are not readily visible when resolved (separated) on an agarose gel. In order to visualize the molecules, a DNA dye must be administered to the gel. In research labs, a DNA intercalating agent called Ethidium Bromide is added to the molten gel and will bind to the DNA of the samples when run. Ethidium Bromide can then be visualized on a UV box that will fluoresce the compound and reveal bands where DNA is accumulated. Since Ethidium Bromide is known as a carcinogen, teaching labs will use a safer DNA intercalating agent known as Sybr Green. This can be visualized in a similar fashion, but will fluoresce a green color instead.

Agarose gels visualized on a UV transilluminator. Left shows a gel with Ethidium bromide. Right shows a gel with Sybr Green.
Agarose gels are made of and bathed in a buffered solution, usually of Tris-Borate-EDTA (TBE) or Tris-Acetate-EDTA (TAE). Regardless of buffer solution, the buffer provides necessary electrolytes for the current to pass through and maintain the pH of the solution.
DNA samples are prepared in a buffer similar to the solution that it will be run in to ensure that the phosphate backbone of the DNA remains deprotonated and moves to the positive electrode. Additionally, glycerol or another compound is added to this buffer in order for the solution to sink into the wells without spreading out. A dye is often included in this loading buffer in order to visualize the loading in the wells and to track the relative progression of gel.