- Describe the role of the promoter, the operator, the structural genes, and the regulator in the prokaryotic chromosome.
- Describe the basic functioning of the lac operon and their differences.
- List the major differences between prokaryotic and eukaryotic gene regulation.
- Define histone, nucleosome, and looped domain.
- Distinguish between euchromatin and heterochromatin, and discuss how they are involved in gene regulation.
- Describe the four levels of regulation of gene expression in Eukaryotes: transcriptional, post-transcriptional, translational and post-translational regulation.
- Define the term mutation, and explain the difference among missense, nonsense, frameshift, and point mutation.
- Distinguish between germ-line and somatic mutations.
- Understand the link between mutation and diseases.
Contents
The Lactose Intolerance of Bacteria
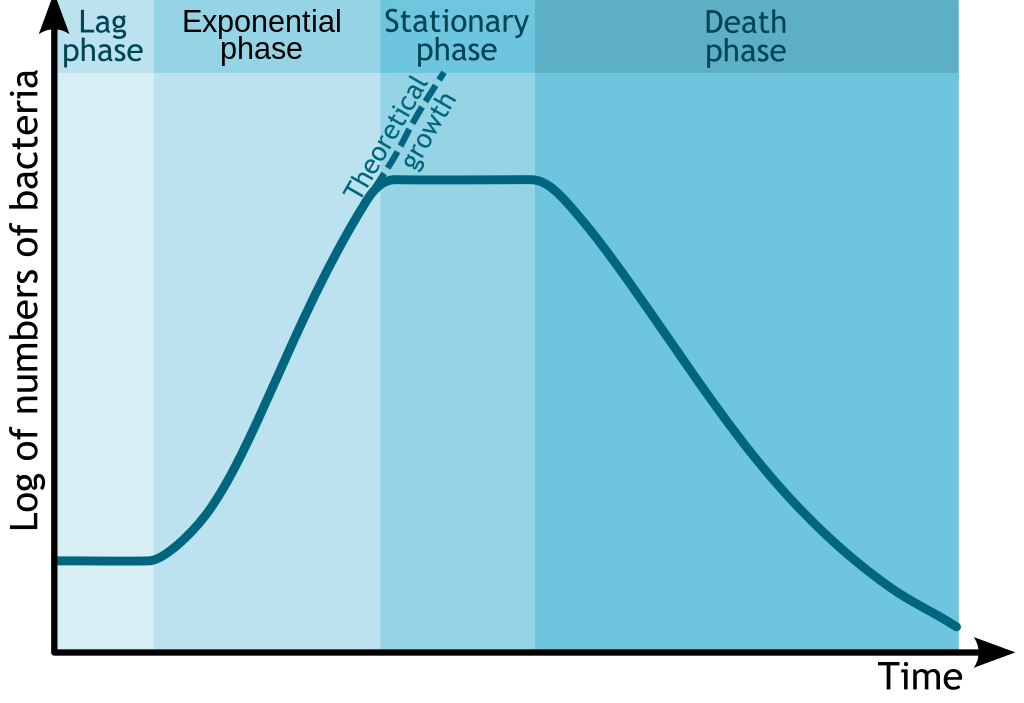
The standard growth kinetics of E. coli are described by the curve. Credit: Michał Komorniczak (CC-BY-SA 3.0)
Glucose is the preferred energy source of cells. François Jacob and Jacques Monod sought to understand how bacteria made decisions to switch between different sugars as sources of energy. Jacob and Monod found that if glucose and lactose were presented as food for bacteria, there would be a biphasic growth pattern.

Jacob and Monod came to understand that the glucose would first be utilized (preferred source) and the lactose would be digested after the depletion of glucose. This occurred because, under normal situations the bacteria would not have access to lactose and would waste energy by creating enzymes to digest it. The enzyme β-galactosidase, which is responsible for digesting lactose to the monomers galactose and glucose would only be induced under the conditions of low glucose and high lactose. Monod found that when lactose was the sole sugar, the expression of the enzyme β-galactosidase was induced and displayed a monophasic growth with a delay.
The Lac Operon
Jacob and Monod later found that the genes involved in utilizing lactose were clustered together in proximity under a coordinated control mechanism. This became known as the Lac operon.

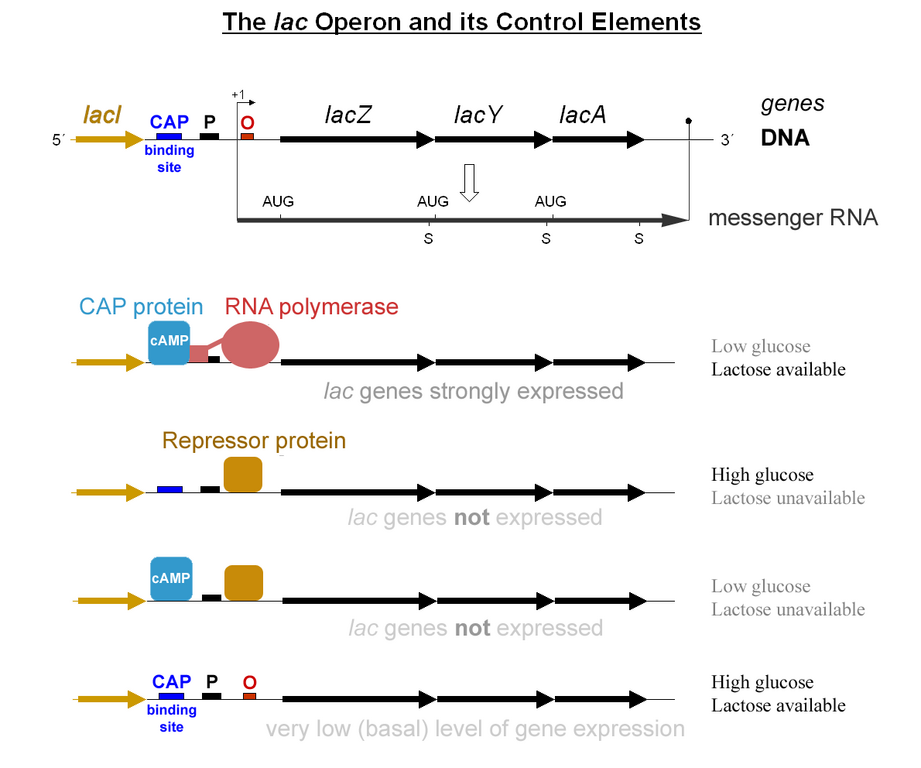
A schematic of the Lac Operon. LacZ, LacY and LacA are transcribed as a single mRNA.
The usage of lactose as a source of energy is preferred by bacteria when glucose is not present. In the presence of abundant glucose, it would be a waste of energy and cellular resources to commit to synthesizing the mRNA and the protein for β-galactosidase. Unless lactose is present, a protein binds to a portion of the Lac promoter referred to as the operator. This repressor protein is encoded by another gene (LacI) outside of the gene cluster. Occasionally, the repressor unbinds to the operator and RNA Polymerase is permitted to transcribe the LacZ gene (β-galactosidase), LacY gene (permease), and LacA gene (acetylase).This “leakiness” of expression is important since the permease protein is needed on the surface of the cell to allow lactose into the cell if it is present in the environment. The presence of lactose causes the repressor to fall off the operator to grant RNA pol access to the DNA. When glucose is low, a protein called CAP (Catabolite Activated Protein), binds to the Lac promoter and works as an recruiter of RNA pol. The coordinated effects of CAP activation and Lac Repressor inactivation yields high transcription of the operon.

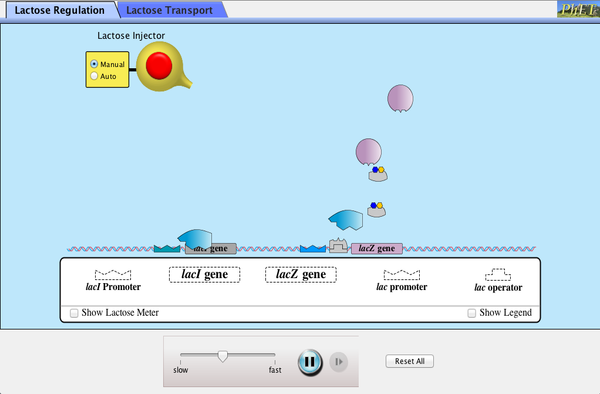
Lac Operon Simulation
Launch the simulation below to explore the coordinated activation of the Lac Operon.