Restriction fragment length polymorphism (RFLP) is a technique that exploits variations in DNA sequences. DNA from differing sources will have variations or polymorphisms throughout the sequence. Using restriction enzymes, these differences in sequences may be teased out. However, if one were to take the entirety of the human genome and chop it up with a restriction enzyme, many indecipherable fragments would be made. In fact, the resulting agarose gel would simply show a large smear of DNA. RFLP analysis requires that a probe to a specific area of DNA be used to identify specific locations. Agarose gels can then be transferred to a membrane or filter where they would be hybridized to these radioactive probes.
RFLP analysis was designed for forensic science to discriminate between people. Since people are 2n, they have pairs of homologous chromosomes with the same loci. However, these loci may contain different alleles. In this case, the phenotype for these alleles is the actual sequence that may or may not contain restriction sites. The presence or absence of a restriction site may arise from single nucleotide polymorphisms (SNPs) that reveal the natural variation between people.
The schematic below illustrates a comparison of restriction profiles between two sources.
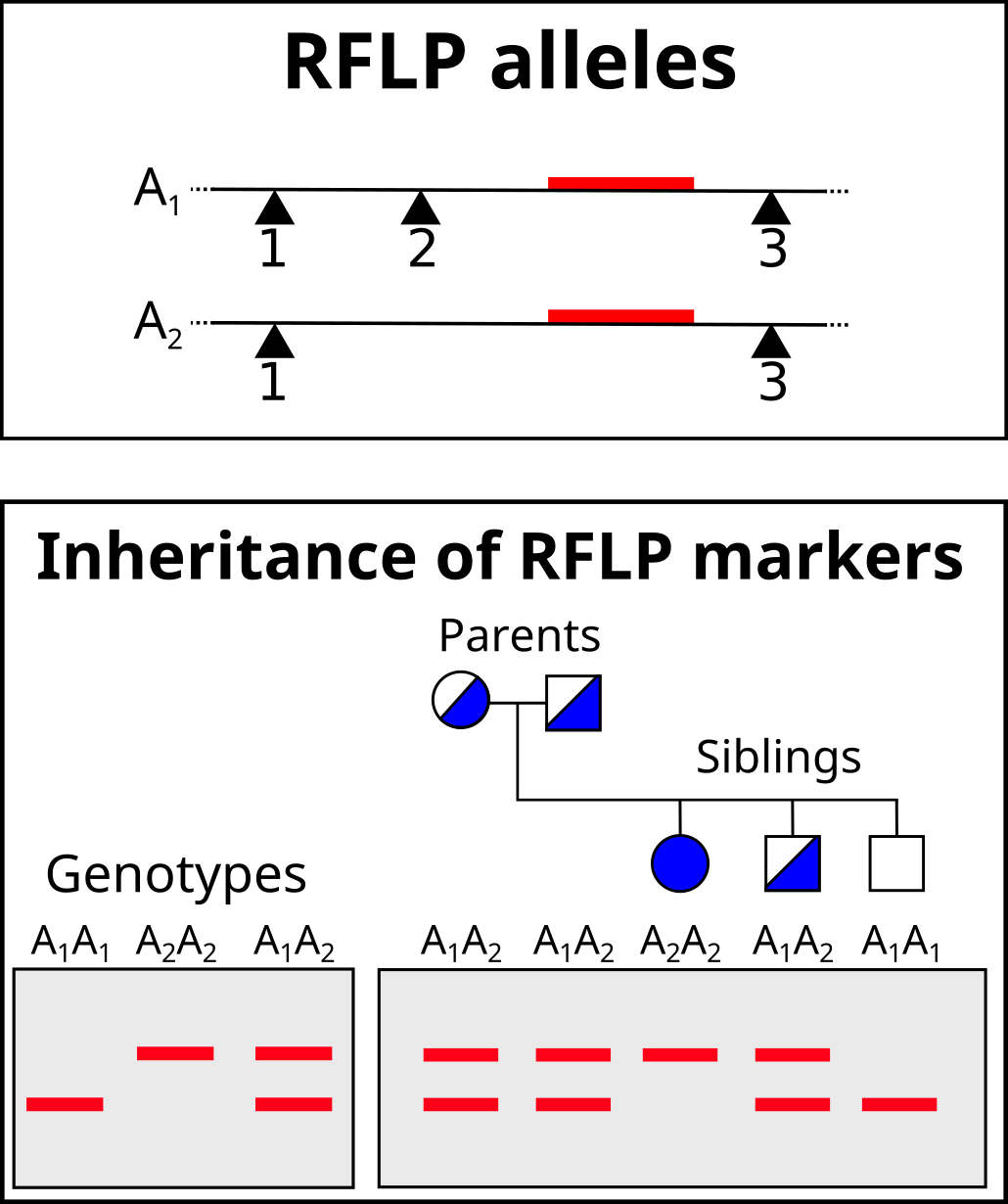
Note that the probe overlaps a restriction site in one of the alleles. This probe will be able to bind to both fragments given sufficient sequence overlap.
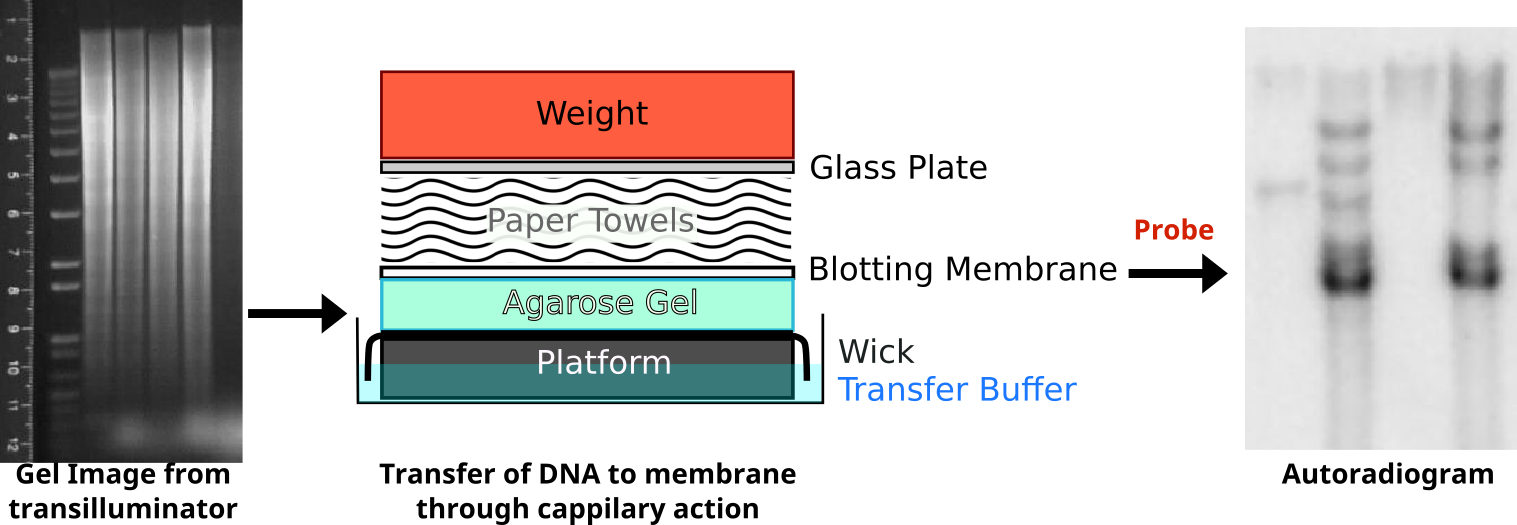
Upon resolving on an agarose gel, genomic DNA that does not hybridize with the probe will obscure the locus of interest as a large smear. A filter is placed on top of the agarose and pressed against it to transfer the DNA in a process called Southern blotting. Following a lengthy transfer, the filter is denatured to and incubated with the radioactive probe. To visualize this probe hybridization, film is exposed to the filter and processed.
RFLPs represent inheritable markers and can reveal relationships between different individuals. A pedigree can illustrate the relationship of the inherited alleles. The technique can be more informative if using multiple probes simultaneously for different loci or to use multi-locus probes that hybridize to multiple locations.
While RFLPs can arise from SNPs, they may also be caused by the expansion or contraction of repeated elements between restriction sites. These repeated elements of DNA are referred to as variable number tandem repeats (VNTR) and illustrate polymorphisms that normally occur in non-coding regions of the genome.
Table of Contents
Explore: DNA Fingerprinting
Click on the image below to learn more about the process of DNA fingerprinting.
Click on the image below to play a simulation of creating a DNA fingerprint.
Print this page





